cdapath:Vol1Profiles: Unterschied zwischen den Versionen
(→10.3 APSR Required Actor Groupings) |
(→10.4.2.1.3 Single Report Text Example) |
||
| Zeile 313: | Zeile 313: | ||
ONCOPROTEIN.<br/> | ONCOPROTEIN.<br/> | ||
HER2/NEU FISH RESULT: NEGATIVE FOR AMPLIFICATION OF HER2/NEU.<br/> | HER2/NEU FISH RESULT: NEGATIVE FOR AMPLIFICATION OF HER2/NEU.<br/> | ||
| + | |||
| + | |||
| + | Procedure steps: | ||
| + | |||
| + | RIGHT BREAST FIVE CORES 8-9:00" (ULTRASOUND GUIDED NEEDLE CORE BIOPSY) PARAFFIN BLOCK NUMBER: A1.:FOUR SECTIONS FOR EACH STAIN: | ||
| + | |||
| + | RIGHT BREAST FIVE CORES 8-9:00 | ||
| + | |||
| + | PARAFFIN BLOCK NUMBER: A1 | ||
| + | |||
| + | slide from block A1 HE stained | ||
| + | |||
| + | slide from block A1 ER Immunohistochemistry | ||
| + | |||
| + | slide from block A1 PR Immunohistochemistry | ||
| + | |||
| + | slide from block A1 EGFR (PharmDX) Immunohistochemistry | ||
| + | |||
| + | slide from block A1 HER2 Immunohistochemistry | ||
| + | |||
| + | slide block from A1 HER2 FISH | ||
====10.4.2.2 Use Case #2: Multi-step Report==== | ====10.4.2.2 Use Case #2: Multi-step Report==== | ||
Aktuelle Version vom 7. Februar 2018, 11:25 Uhr
Inhaltsverzeichnis
- 1 VOLUME 1 - PROFILES
- 1.1 10 Anatomic Pathology Structured Report (APSR) Profile
- 1.2 10.1 APSR Actors/Transactions
- 1.3 10.2 APSR Actor Options
- 1.4 10.3 APSR Required Actor Groupings
- 1.5 10.4 APSR Overview
- 1.6 10.5 APSR Security Considerations
- 1.7 10.6 APSR Cross Profile Considerations
VOLUME 1 - PROFILES
10 Anatomic Pathology Structured Report (APSR) Profile
This content profile describes an anatomic pathology structured report (APSR) as a digital document to be shared or exchanged between pathology laboratories and other care providers and institutions.
Anatomic pathology structured reports document the findings on specimens removed from patients for diagnostic or therapeutic reasons. This information can be used for patient care, clinical research and epidemiology. Standardizing and computerizing anatomic pathology reports is necessary to improve the quality of reporting and to facilitate the exchange and reuse of the content of these reports.
This content profile describes a digital anatomic pathology report shared in a human-readable format, which may include images, and which also contains findings and observations in a machine-readable format, to facilitate the integration of these into the database of a consumer system, and to enable the application of automated reasoning to this content.
The scope of this IHE content profile covers all fields of anatomic pathology (cancers, benign neoplasms as well as non-neoplastic conditions) as well as cytopathology.
Goldsmith J.D. et al.[1] is the first source of specification for this content profile. This article delineates the required, preferred, and optional elements which should be included in any report of surgical pathology.
This source is complemented by the “Cancer Checklists” produced by the College of American Pathologists[2], and by the “comptes rendus d’anatomopathologie : données minimales à renseigner pour une tumeur primitive” produced by the French society of pathology (SFP[3]) for the French cancer institute (INCa [4]), and by the German "Guideline Pathology / Neuropathology" (formerly TM-30) of the Sector Committee Pathology for the implementation of DIN EN ISO/EC 17020.
This profile has also benefited from the guidance on cancer AP reports provided by the North-American Association of Central Cancer Registries; some of the example snippets captured in the profile leverage the NAACCR Standards for Cancer Registries, Volume V, Pathology Laboratory Electronic Reporting.
10.1 APSR Actors/Transactions
This section defines the actors, transactions, and/or content modules in this profile. General definitions of actors are given in the Technical Frameworks General Introduction Appendix A published here.
Figure 10.1-1 shows the actors directly involved in the APSR Profile and the direction that the content is exchanged.
A product implementation using this profile must group actors from this profile with actors from a workflow or transport profile to be functional. The grouping of the content module described in this profile to specific actors is described in more detail in the “Required Actor Groupings” section below.
Figure 10.1-1 APSR Actor Diagram
Table 10.1-1 lists the content module(s) defined in the APSR profile. To claim support with this profile, an actor shall support all required content modules (labeled “R”) and may support optional content modules (labeled “O”).
Table 10.1-1: <Profile Acronym>Profile - Actors and Content Modules
| Actors | Content Modules | Optionality | Reference |
|---|---|---|---|
| Content
Creator |
Anatomic Pathology Structured Report 1.3.6.1.4.1.19376.1.8.1.1.1 |
R | PaLM TF-3: 6.3.1.2 |
| Content
Consumer |
Anatomic Pathology Structured Report 1.3.6.1.4.1.19376.1.8.1.1.1 |
R | PaLM TF-3: 6.3.1.2 |
10.1.1 Actor Descriptions and Actor Profile Requirements
Most requirements are documented in Content Modules (Volume 3). This section documents any additional requirements on profile’s actors.
10.2 APSR Actor Options
Options that may be selected for each actor in this profile are listed in the table 10.2-1. These options are further detailed in PCC Technical Framework Volume 2 as indicated in the rightmost column.
Table 10.2-1 Anatomic Pathology Structured Report - Actors and Options
| Actor | Option Name | Reference |
|---|---|---|
| Content
Creator |
None | |
| Content
Consumer |
View Option (1)
Document Import Option (1) Section Import Option (1) |
PCC TF-2:3.1.1
PCC TF-2:3.1.2 PCC TF-2:3.1.3 |
Note 1: The Content Consumer Actor shall support at least one of these options.
10.3 APSR Required Actor Groupings
An Actor from this profile (Column 1) shall implement all of the required transactions and/or content modules in this profile in addition to all of the transactions required for the grouped actor (Column 2).
In some cases, required groupings are defined as at least one of an enumerated set of possible actors; this is designated by merging column one into a single cell spanning multiple potential grouped actors. Notes are used to highlight this situation.
Section 10.5 describes some optional groupings that may be of interest for security considerations and section 10.6 describes some optional groupings in other related profiles.
Table 10.3-1: Anatomic Pathology Structured Report - Required Actor Groupings
| APSR Actor | Actor to be grouped with | Reference | Content Bindings Reference |
|---|---|---|---|
| Content
Creator |
ITI XDS.b Document Source OR ITI XDM Portable Media Creator OR ITI XDR Document Source OR ITI MHD Document Source |
ITI TF-1:10
|
|
| Content
Consumer |
ITI XDS.b Document Consumer OR ITI XDM Portable Media Consumer OR ITI XDR Document Recipient OR ITI MHD Document Consumer |
ITI TF-1:10
ITI TF-1:16
ITI TF-1:15
ITI TF-1:33 |
Note 1: Each actor of APSR SHALL be grouped with at least one of the ITI actors listed in its table row.
10.4 APSR Overview
10.4.1 Concepts
This content profile represents a common digital document model applicable to any structured report for surgical pathology in all fields of anatomic pathology (cancers, benign neoplasms, non-neoplastic conditions) as well as for cytopathology.
This common model is composed of a header conveying the context of care (patient, care providers, pathologists, laboratories, order, act documented …) and a body. The body organizes the human-readable content of the report in a number of sections. Each section may also provide machine-readable content in a repeatable “entry” embedded in the section. This common model defines the order of appearance, cardinalities and internal structure of each section, and of each entry embedded in each section.
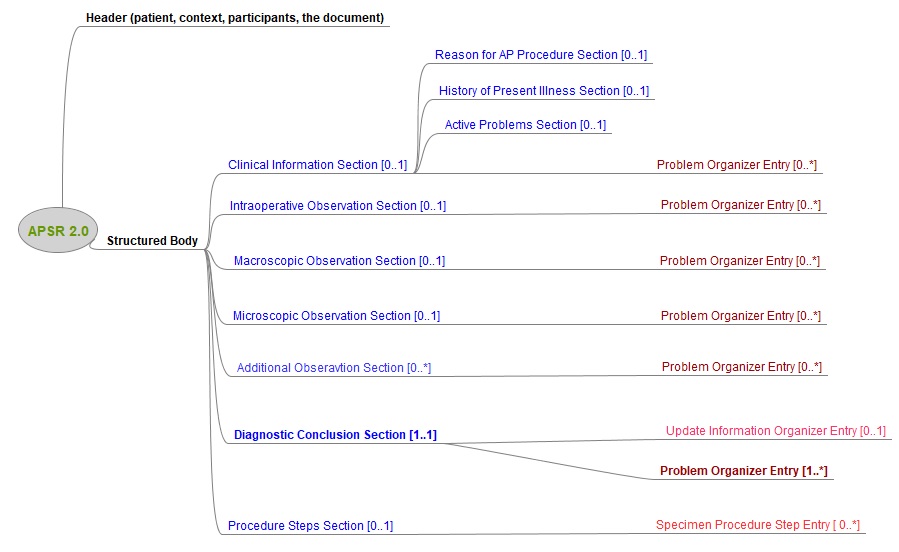
Figure 10.4.1-1 shows this general model applicable to any pathology digital report.
Figure 10.4.1-1 Common model for a digital anatomic pathology structured report
Note 1: The only section that is mandatory is the Diagnostic Conclusion section.
10.4.2 Use Cases
10.4.2.1 Use case #1: Single Report
Anatomic pathology order fulfilled by a pathology laboratory produces a report.
10.4.2.1.1 Single Report Use Case Description
Dr. Eva Surgeon, PhD, takes a ultrasound guided core biopsy from a breast tumor from patient Eve Onewoman, born on Sept 21 1971, requests the procedure “breast core biopsy specimen - pathological examination” and sends the specimen to the anatomic pathology laboratory of the Cancer Institute. One specimen (five cores) is accessioned by the anatomic pathology laboratory under the accession number A710240008. The staff performs a macroscopic examination of the specimen; gross imaging is performed if needed. The specimen with the specimen ID A710240008_A is processed for microscopic examination and other special ancillary techniques or tissue banking if needed. During the imaging interpretation process, microscopic imaging is performed if needed. At the end of the interpretation process of the macroscopic and microscopic observations and some ancillary techniques, done by the pathologists Dr. Marcel Pathologist, PhD, and Dr. Jonas Jones, M.D., Dr. Marcel Pathologist queries the Content Creator application for the appropriate APSR template, fills the form, binds some relevant images and/or regions of interest to specific observations, validates and signs the digital report.
10.4.2.1.2 Single Report Process Flow
Intentionally left blank
10.4.2.1.3 Single Report Text Example
Macroscopic observation
A. "RIGHT BREAST FIVE CORES 8-9:00" (ULTRASOUND GUIDED NEEDLE CORE BIOPSY):
Microscopic observation
INVASIVE ADENOCARCINOMA OF THE BREAST.
ICD-O-3-CLASSIFICATION: C50.3 M8500/33
HISTOLOGIC TYPE: DUCTAL.
NOTTINGHAM COMBINED HISTOLOGIC GRADE: 1 OF 3.
TUBULE FORMATION SCORE: 2.
NUCLEAR PLEOMORPHISM SCORE: 2.
MITOTIC RATE SCORE: 1.
IN-SITU CARCINOMA: EQUIVOCAL.
BREAST CANCER BIOMARKER STUDIES:
PARAFFIN BLOCK NUMBER: A1.
ER INTERPRETATION: POSITIVE ESTROGEN RECEPTOR ACTIVITY (ALLRED SCORE = 8, Percentage of positive cells = 85%, Staining Intensity score = 3).
PR INTERPRETATION: POSITIVE PROGESTERONE RECEPTOR ACTIVITY (ALLRED SCORE = 8).
DAKO EGFR PHARMDX IMMUNOHISTOCHEMISTRY: NEGATIVE (0) FOR EXPRESSION OF
EPIDERMAL GROWTH FACTOR RECEPTOR.
HER2/NEU IMMUNOHISTOCHEMISTRY: AMBIGUOUS(2+) FOR OVEREXPRESSION OF HER2/NEU
ONCOPROTEIN.
HER2/NEU FISH RESULT: NEGATIVE FOR AMPLIFICATION OF HER2/NEU.
Diagnostic conclusion
A. "RIGHT BREAST FIVE CORES 8-9:00" (ULTRASOUND GUIDED NEEDLE CORE BIOPSY):
INVASIVE ADENOCARCINOMA OF THE BREAST.
ICD-O-3-CLASSIFICATION: C50.3 M8500/33
HISTOLOGIC TYPE: DUCTAL.
NOTTINGHAM COMBINED HISTOLOGIC GRADE: 1 OF 3.
TUBULE FORMATION SCORE: 2.
NUCLEAR PLEOMORPHISM SCORE: 2.
MITOTIC RATE SCORE: 1.
IN-SITU CARCINOMA: EQUIVOCAL.
BREAST CANCER BIOMARKER STUDIES:
PARAFFIN BLOCK NUMBER: A1.
ER INTERPRETATION: POSITIVE ESTROGEN RECEPTOR ACTIVITY (ALLRED SCORE = 8, Percentage of positive cells = 85%, Staining Intensity score = 3).
PR INTERPRETATION: POSITIVE PROGESTERONE RECEPTOR ACTIVITY (ALLRED SCORE = 8).
DAKO EGFR PHARMDX IMMUNOHISTOCHEMISTRY: NEGATIVE (0) FOR EXPRESSION OF
EPIDERMAL GROWTH FACTOR RECEPTOR.
HER2/NEU IMMUNOHISTOCHEMISTRY: AMBIGUOUS(2+) FOR OVEREXPRESSION OF HER2/NEU
ONCOPROTEIN.
HER2/NEU FISH RESULT: NEGATIVE FOR AMPLIFICATION OF HER2/NEU.
Procedure steps:
RIGHT BREAST FIVE CORES 8-9:00" (ULTRASOUND GUIDED NEEDLE CORE BIOPSY) PARAFFIN BLOCK NUMBER: A1.:FOUR SECTIONS FOR EACH STAIN:
RIGHT BREAST FIVE CORES 8-9:00
PARAFFIN BLOCK NUMBER: A1
slide from block A1 HE stained
slide from block A1 ER Immunohistochemistry
slide from block A1 PR Immunohistochemistry
slide from block A1 EGFR (PharmDX) Immunohistochemistry
slide from block A1 HER2 Immunohistochemistry
slide block from A1 HER2 FISH
10.4.2.2 Use Case #2: Multi-step Report
Reporting includes multiple successive steps.
10.4.2.2.1 Multi-step Report Use Case Description
A surgeon removes a breast tumor from a patient, requests the procedure “breast surgical specimen - frozen sections & pathological examination”, and “breast surgical specimen - pathological examination” and sends the specimen(s) to the anatomic pathology laboratory.
Specimens are accessioned by the anatomic pathology department. The staff performs a macroscopic examination of the specimens, gross imaging is performed if needed. The specimens are processed for intraoperative observation if needed, and tissue banking if needed (e.g., for research purpose). During the imaging interpretation process of frozen sections, microscopic imaging is performed if needed. At the end of the interpretation process, the pathologist queries the Content Creator for the appropriate APSR template, fills the intraoperative observation section, binds some relevant images and/or regions of interest to specific observation(s) if needed, validates and signs (i.e., legally authenticates) the preliminary APSR.
The day after, the specimen(s) are processed for microscopic examination and other special ancillary techniques if needed. During the imaging interpretation process, microscopic imaging is performed if needed. At the end of the interpretation process, pathologist queries the Content Creator for the preliminary APSR, fills the form, binds some relevant images and/or regions of interest to specific observation(s), validates and signs (i.e., legally authenticates) the final APSR.
10.4.2.2.2 Multi-step Report Process Flow
Intentionally left blank
10.5 APSR Security Considerations
See Appendix A of PaLM TF-1.
10.6 APSR Cross Profile Considerations
Intentionally left blank
- ↑ Goldsmith, J.D., et al., “Reporting guidelines for clinical laboratory reports in surgical pathology” Arch Pathol Lab Med, 2008. 132(10): p. 1608-16
- ↑ http://www.cap.org/web/oracle/webcenter/portalapp/pagehierarchy/cancer_protocol_templates.jspx?_adf.ctrl-state=e1f4ltym4_4&_afrLoop=275425457234715#!
- ↑ http://www.sfpathol.org
- ↑ http://www.e-cancer.fr